-
Products
- Respiratory Pathogens Nucleic Acid Detection
- Six Respiratory Viruses Nucleic Acid Test Kit (PCR-Melting Curve Method)
- Respiratory System Infection Pathogen Targeted Next-generation Sequencing (tNGS) Kit
- Bloodstream Infection Pathogen Targeted Next-generation Sequencing (tNGS) Kit
- Central Nervous System Infection Pathogen Targeted Next-generation Sequencing (tNGS) Kit
- Genitourinary Tract Infection Pathogen Targeted Next-generation Sequencing (tNGS) Kit
- MAPMI Ultra-Broad-Spectrum Pathogen mNGS Test
- Mycobacteria Nucleic Acid Detection
- Mycobacterium Tuberculosis Rifampicin Resistance Detection Kit (PCR-Melting Curve Method)
- Mycobacterium Tuberculosis Isoniazid Resistance Detection Kit (PCR-Melting Curve Method)
- Mycobacteria Species Identification
- MTB Drug Resistant Detection
- Human Papillomavirus (HPV) Genotyping Detection
- Human Papillomavirus (HPV) Nucleic Acid Detection Kit (PCR - Fluorescent Probe Method)
- Neisseria Gonorrhoea (NG) Nucleic Acid Detection Kit (PCR-Fluorescent Probe Method)
- Ureaplasma Urealyticum (UU) Nucleic Acid Detection Kit (PCR-Fluorescent Probe Method)
- Monkeypox Virus (MPXV) Nucleic Acid Detection Kit (PCR-Fluorescent Probe Method)
- Preimplantation Embryo Genetic Screening (PGS)
- Drug-Related Deafness Gene Mutations Detection
- 100 Loci of 18 Deafness-Related Genes Detection
- Hereditary Deafness Gene Detection (227 Genes+Mitochondrial Genome)
- Fetal Aneuploidies (Trisomy 21, Trisomy 18 and Trisomy 13) Detection
- Fetal Chromosomal Aneuploidy and Microdeletions/ Microduplications Detection (NIPT-Plus)
- CNV-seq Detection (Semiconductor Sequencing)
- Karyotype Analysis
- CMA Detection
- Folic Acid Utilization Ability Genetic Detection
- Y Chromosome Microdeletion Gene Detection
- Tandem Mass Spectrometry Screening for Genetic Metabolic Diseases
- DMD (Duchenne muscular dystrophy)/ BMD (Becker muscular dystrophy) Genetic Detection
- 17 Common Single-gene Disorders
- Neuromuscular Disorders
- Epilepsy-related Gene Detection
- GJB2 Gene Coding Region and Splicing Site Detection
- Human Mitochondrial Genome Detection
- Clinical WES
- Individual WES
- WES+CNV-seq
- Precise Diagnosis and Treatment for Brain Cancer
- Targeted Medication Guidance for Lung Cancer
- Targeted Medication Guidance for Colorectal Cancer
- Individualized Medication Guidance for Gastrointestinal Cancer
- Genetic Testing for BRCA1/2 Targeted-Drug Guidance (Germline + Somatic)
- Individualized Medication Guidance for Breast Cancer
- Individualized Medication Guidance for Gynecologic Cancer
- Breast Cancer Prognosis Assessment
- Genetic Testing for Hereditary Breast/Ovarian Cancer(BRCA1/2)
- Individualized Medication Guidance for Prostate Cancer
- Genetic Testing for Blood Cancer
- Individualized Chemotherapy Drugs For Cancers (10 Genes)
- Individualized Medication Guidance for Cancer (800 Genes)
- The Disease Susceptibility Genetic Test-Lung Cancer
- The Disease Susceptibility Genetic Test-Liver Cancer
- The Disease Susceptibility Genetic Test-Stomach Cancer
- The Disease Susceptibility Genetic Test-Breast Cancer
- The Disease Susceptibility Genetic Test-Colorectal Cancer
- The Disease Susceptibility Genetic Test-Esophageal Cancer
- The Disease Susceptibility Genetic Test-High Incidence Cancer (6 Items)
- The Disease Susceptibility Genetic Test-High Incidence Cancer (10 Items)
- The Disease Susceptibility Genetic Test-Cancer Screening Package (29 Items)
- The Disease Susceptibility Genetic Test-Anti-Cancer Package A
- The Disease Susceptibility Genetic Test-Anti-Cancer Package C
- The Disease Susceptibility Genetic Test-Alzheimer's Disease
- The Disease Susceptibility Genetic Test-Thrombotic Disorders (5 Items)
- The Disease Susceptibility Genetic Test-Diabetes (4 Items)
- The Disease Susceptibility Genetic Test-Cardiovascular Disease (14 Items)
- The Disease Susceptibility Genetic Test-Chronic Disease Package (34 Items)
- The Disease Susceptibility Genetic Test-Chronic Disease Package (63 Items)
- The Disease Susceptibility Genetic Test-APOE
- The Disease Susceptibility Genetic Test-ADIPO
- The Disease Susceptibility Genetic Test-Full Package (150 Items)
- The Disease Susceptibility Genetic Test-Obesity
- The Disease Susceptibility Genetic Test-High Myopia
- Metabolic Testing-Alcohol Metabolism Test
- Metabolic Testing-Tobacco Damage
- Metabolic Testing-Alcohol Damage
- Metabolic Testing-Food Intolerance (50 Items)
- Metabolic Testing-Food Intolerance (101 Items)
- Intestinal Flora Testing
-
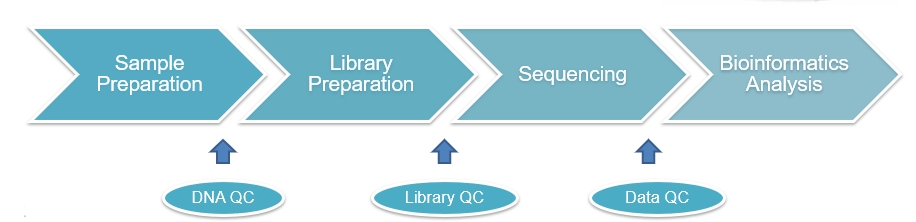
Sequencing Service
- Agricultural SNP Microarray
- Illumina 850K Methylation Microarray DNA Methylation Sequencing
- Human SNP Microarray
- Resources
- About CapitalBiotech
- Contact Us
 EN
EN